You identified a compound with the desired effect ?
You are looking for its target(s) ?
Which additional pathways are affected ?
Which proteins are altered in a diseased tissue ?
Get the whole picture of protein expression in multiple pathways !
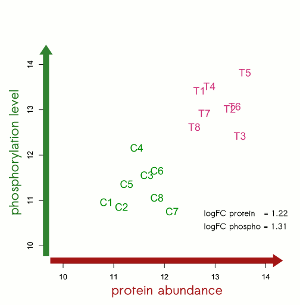
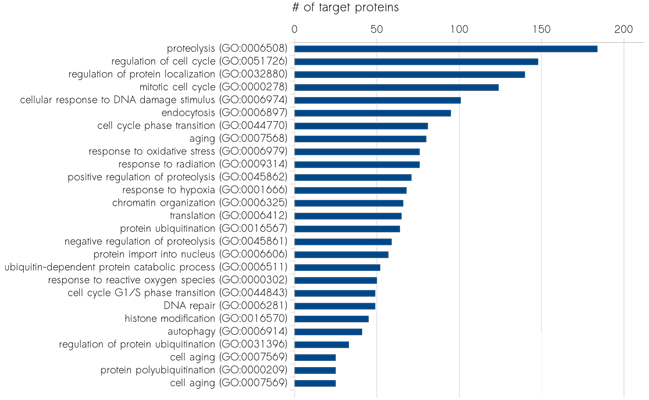
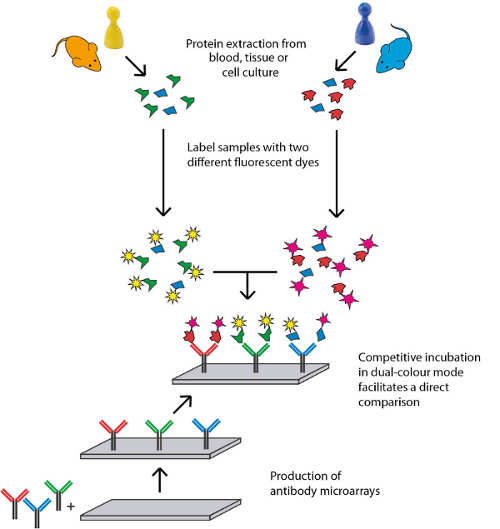
Profound knowledge about protein expression levels is of utmost importance for the discovery and evaluation of new drug targets. Our Scio-Discover Service is fully immuno-based and covers proteins involved in major disease and signalling pathways as well as secreted proteins. For the highly-parallel protein analysis only minimal amounts of sample are required. No sample fractionation or depletion is needed. Our platform is able to process most biological samples without the need for laborious sample preparation, saving time and resources.
Applications:
- Drug Target Identification and verification studies
- Mechanistic studies for various indications
- Correlate Next-Gen-Sequencing Data to protein expression levels
- Profile cytokine and inflammatory status of your samples
Key advantages:
- Low sample consumption
- Investigation of human, murine and rat samples*
- Fast turnaround time, as low as two weeks**
- No need for laborious depletion or fractionation of your samples
- High sensitivity (as sensitive as an ELISA or better)
- High reproducibility (CV < 10%)
- Various sample types can be investigated
- Serum / Plasma
- Tissue
- Cell culture
- Cerebrospinal fluid (CSF)
Service Features:
- Study Layout
- Protein Extraction and Purification
- Protein Quantification
- Protein labelling with high-performance fluorescent dyes
- Automated array incubation and handling
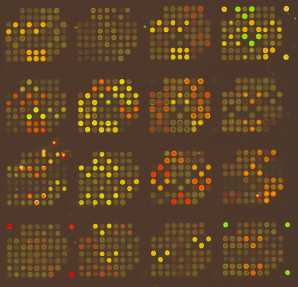
- Array scanning and data acquisition
- Data analysis according to your requirements
- Individual study report
Overview
Scio-Discover Array
Content Biomarker
Discovery Contact
us
* Antibody array is designed to target human proteins. Due to a high sequence homology of vast majority of proteins it was often successfully applied to study murine, rat and ovine models as well [Thorenz A. et al. (2018), Peiro J. et al. (2018), Reichman H. et al. (2019), Raquel F. et al. (2020)].
** depending on study design and sample numbers