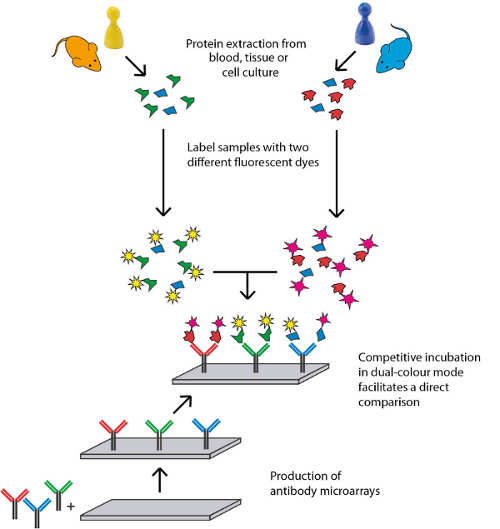
Our scioDiscover and scioPhospho services are based on complex Antibody Microarrays, also called analytical Protein Arrays or Protein Capture Arrays. Within a single experiment 1030 proteins can be profiled in parallel requiring only minimal amounts of sample. The targets proteins were carefully selected over a period of more than 15 years to cover a broad set of different protein classes as well as pathways important in biomedical studies.
Thereby, the protein profiling covers key pathways in various diseases such as cancer, neurological disorders as well as organ failure. Given the wide spectrum of applications and indications the service is ideal for high-content protein expression level analysis from various biological samples. The broad coverage of many important pathways, extracellular and membrane proteins as well as transcription factors, renders scioDiscover a versatile and efficient tool for discovery projects.
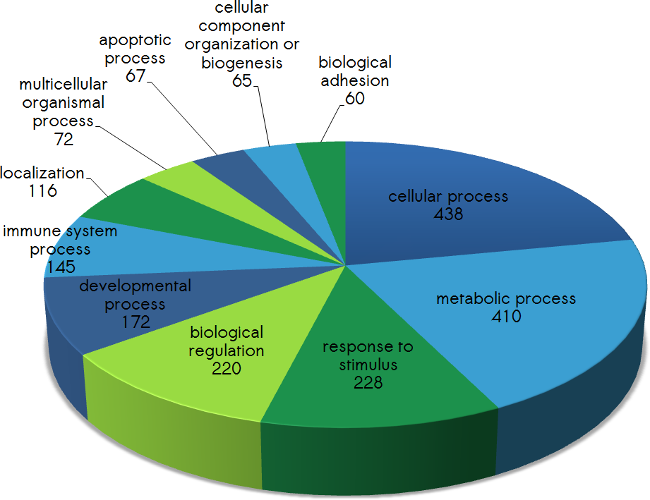
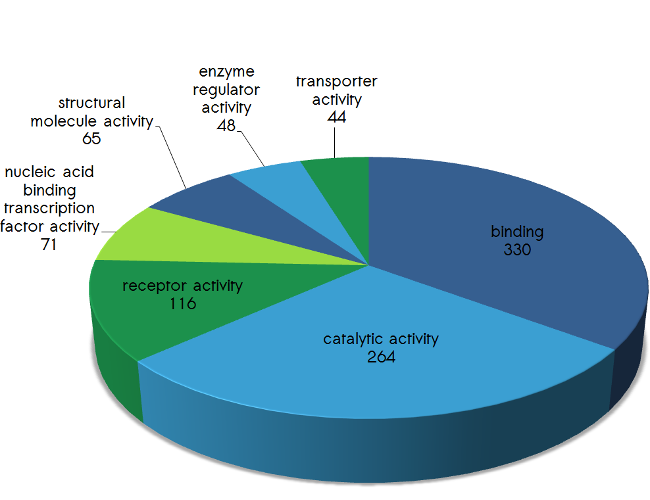
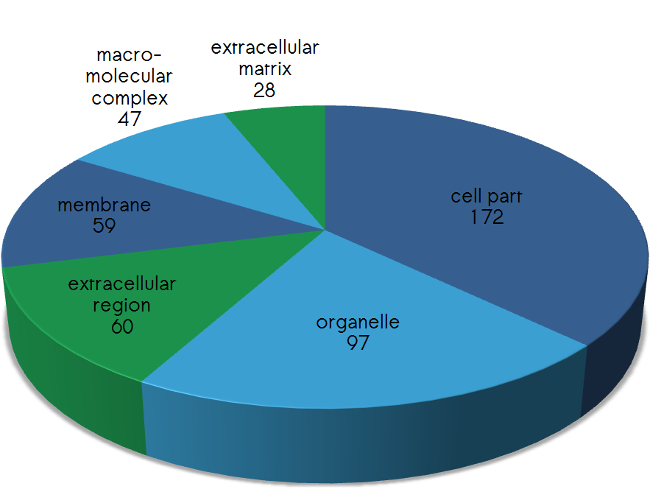
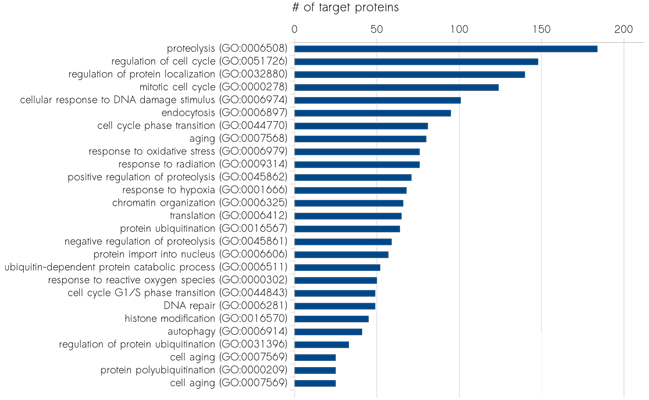
Please find below the distribution of the proteins targeted by scioDiscover according their biological processes (Fig.1), their protein classes (Fig.2), their molecular function (Fig.3) as well as their cellular compartment (Fig.4). In addition, pathways covered by scioDiscover are listed in Table 1.
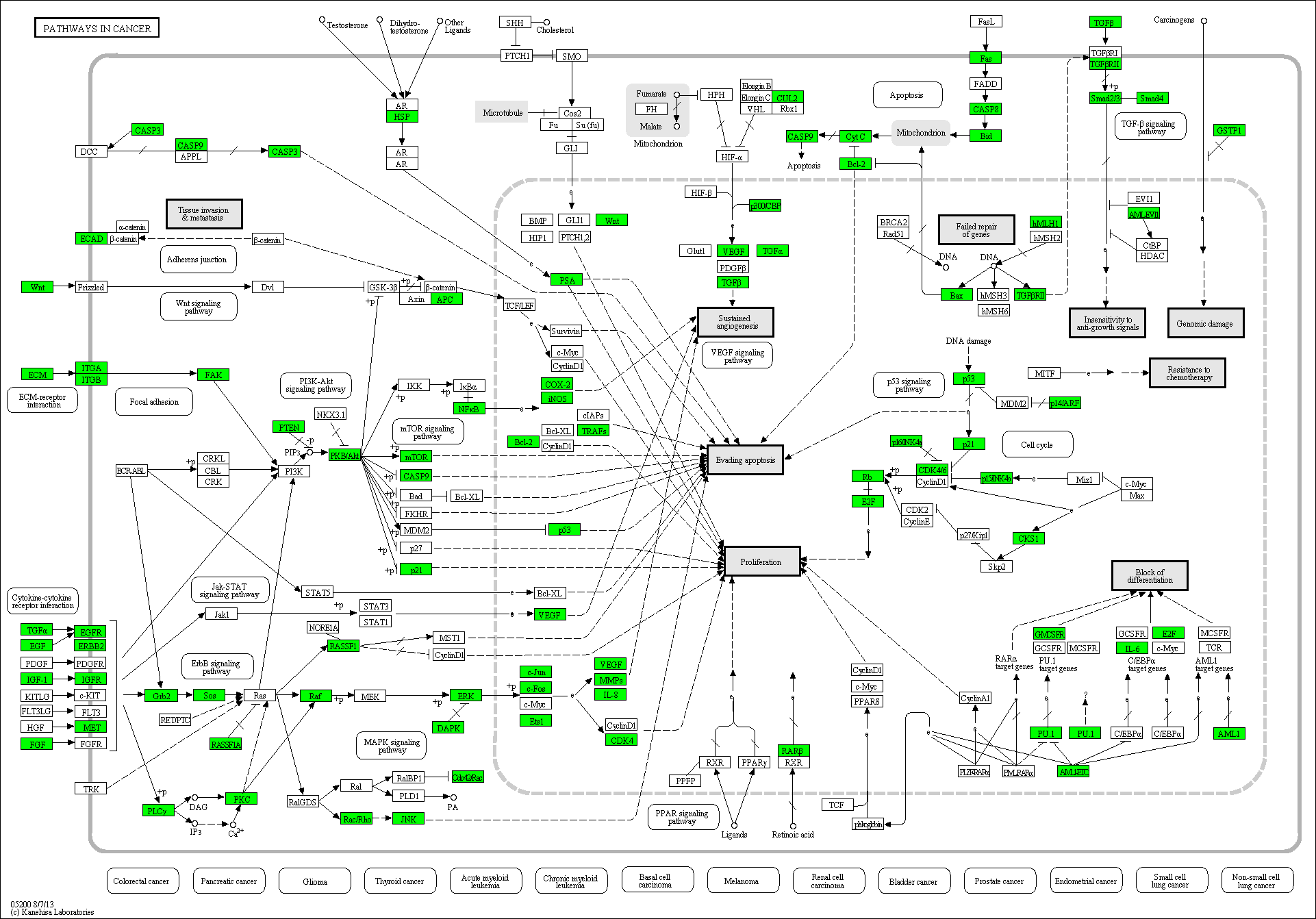
For the pathways TNF signaling, Cell cycle regulation and pathways important in Cancer, Tuberculosis and Huntington's Disease covered targets are depicted in green.
Please contact us for more information regarding targets and pathways covered or to discuss how we can support your study with a highly-parallel protein analysis.
Case study - Acute Kidney Injury
Overview
Scio-Discover Biomarker
Discovery Target
Discovery Contact
us
Classification of target proteins
Coverage of pathways
| Selected Pathways | no. targets |
|---|---|
| Inflammation mediated by chemokine and cytokine signaling (P00031) | 40 |
| Apoptosis signaling (P00006) | 38 |
| Gonadotropin releasing hormone receptor (P06664) | 35 |
| CCKR signaling map (P06959) | 35 |
| Integrin signalling (P00034) | 29 |
| Angiogenesis (P00005) | 28 |
| Interleukin signaling (P00036) | 27 |
| Huntington disease (P00029) | 26 |
| p53 pathway (P00059) | 22 |
| T cell activation (P00053) | 22 |
| B cell activation (P00010) | 21 |
| Ras Pathway (P04393) | 18 |
| Wnt signaling (P00057) | 17 |
| TGF-beta signaling (P00052) | 17 |
| EGF receptor signaling (P00018) | 17 |
| Alzheimer disease-presenilin (P00004) | 15 |
| FAS signaling (P00020) | 15 |
| FGF signaling (P00021) | 14 |
| VEGF signaling (P00056) | 13 |
| Toll receptor signaling (P00054) | 13 |
| Parkinson disease (P00049) | 12 |
| PDGF signaling (P00047) | 12 |
| Insulin / IGF pathway-mitogen activated protein kinase kinase / MAP kinase cascade (P00032) | 12 |
| Alzheimer disease - amyloid secretase (P00003) | 11 |
| p53 pathway feedback loops 2 (P04398) | 10 |
| Endothelin signaling (P00019) | 10 |
| Blood coagulation (P00011) | 10 |
| Plasminogen activating cascade (P00050) | 9 |
| Cytoskeletal regulation by Rho GTPase (P00016) | 9 |
| Interferon-gamma signaling (P00035) | 8 |
| Insulin/IGF pathway-protein kinase B signaling cascade (P00033) | 8 |
| Ubiquitin proteasome (P00060) | 7 |
| Transcription regulation by bZIP transcription factor (P00055) | 7 |
| PI3 kinase (P00048) | 7 |
| Oxidative stress response (P00046) | 7 |
| Cadherin signaling (P00012) | 7 |
Highly parallel protein analysis of TNF Signaling
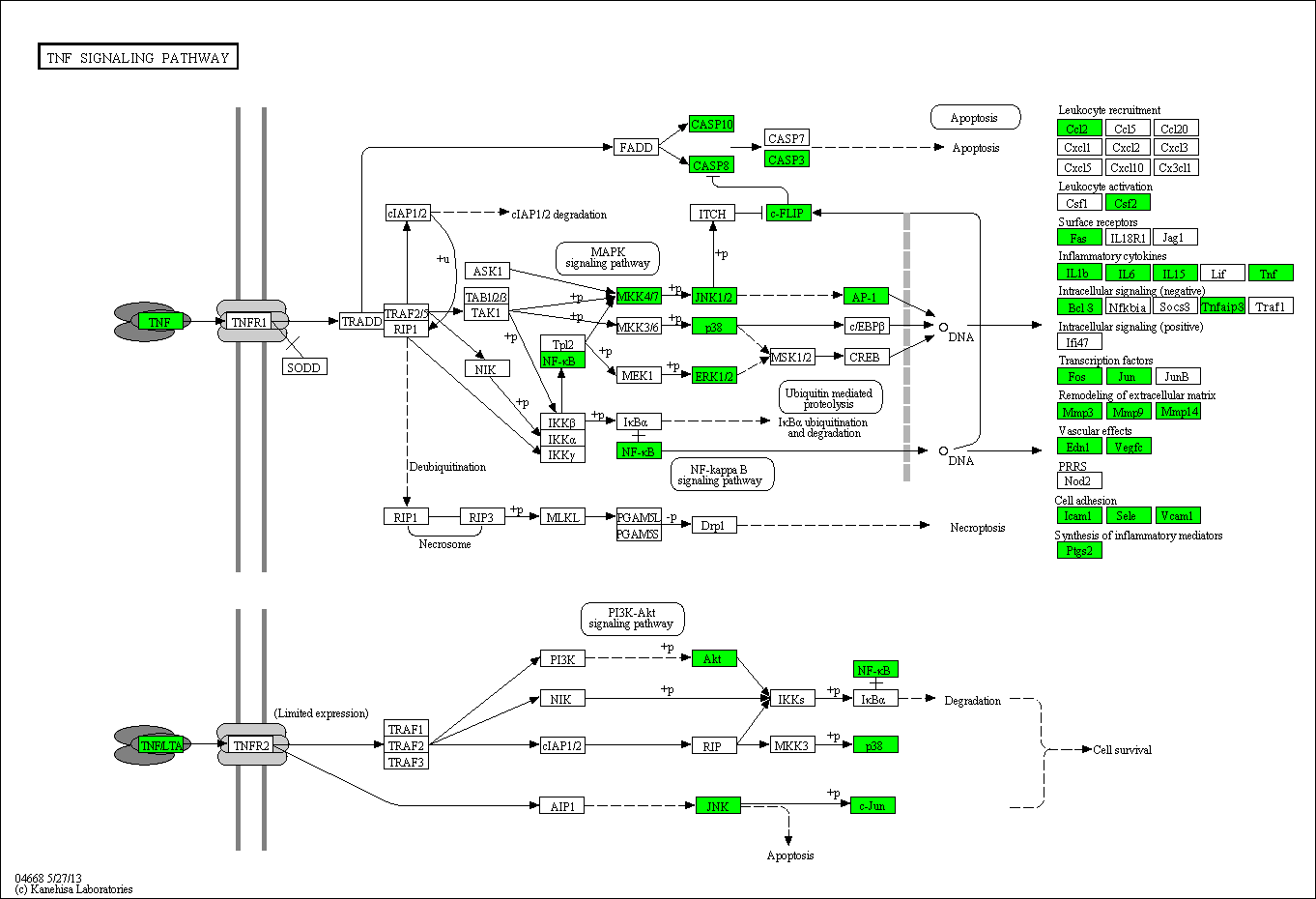
Fig. 5: Proteins in the TNF signalling pathway covered by our protein array are shown in green.
Parallel Analysis of Cell Cycle Proteins by complex antibody arrays
Fig. 6: Cell Cycle Proteins covered by antibody array are depicted in green.
Cancer protein profiling
Fig 7: scioDiscover has a broad coverage of diverse pathways involved in cancer. Proteins in green can be profiled with scioDiscover antibody arrays.
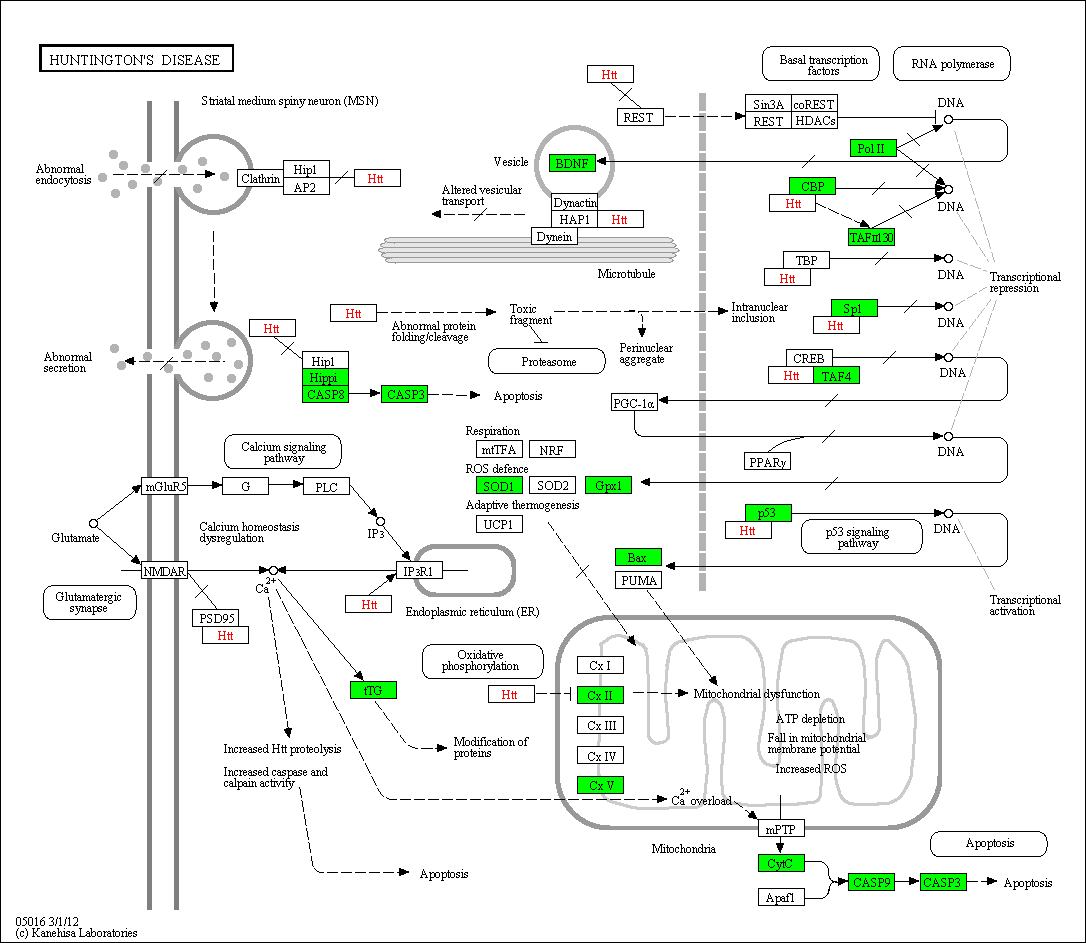
Protein analysis in Huntington's disease
Fig. 8: scioDiscover has a good coverage of proteins involved in various diseases such as Huntington's disease. Proteins analysed in parallel by scioDiscover protein arrays are shown in green
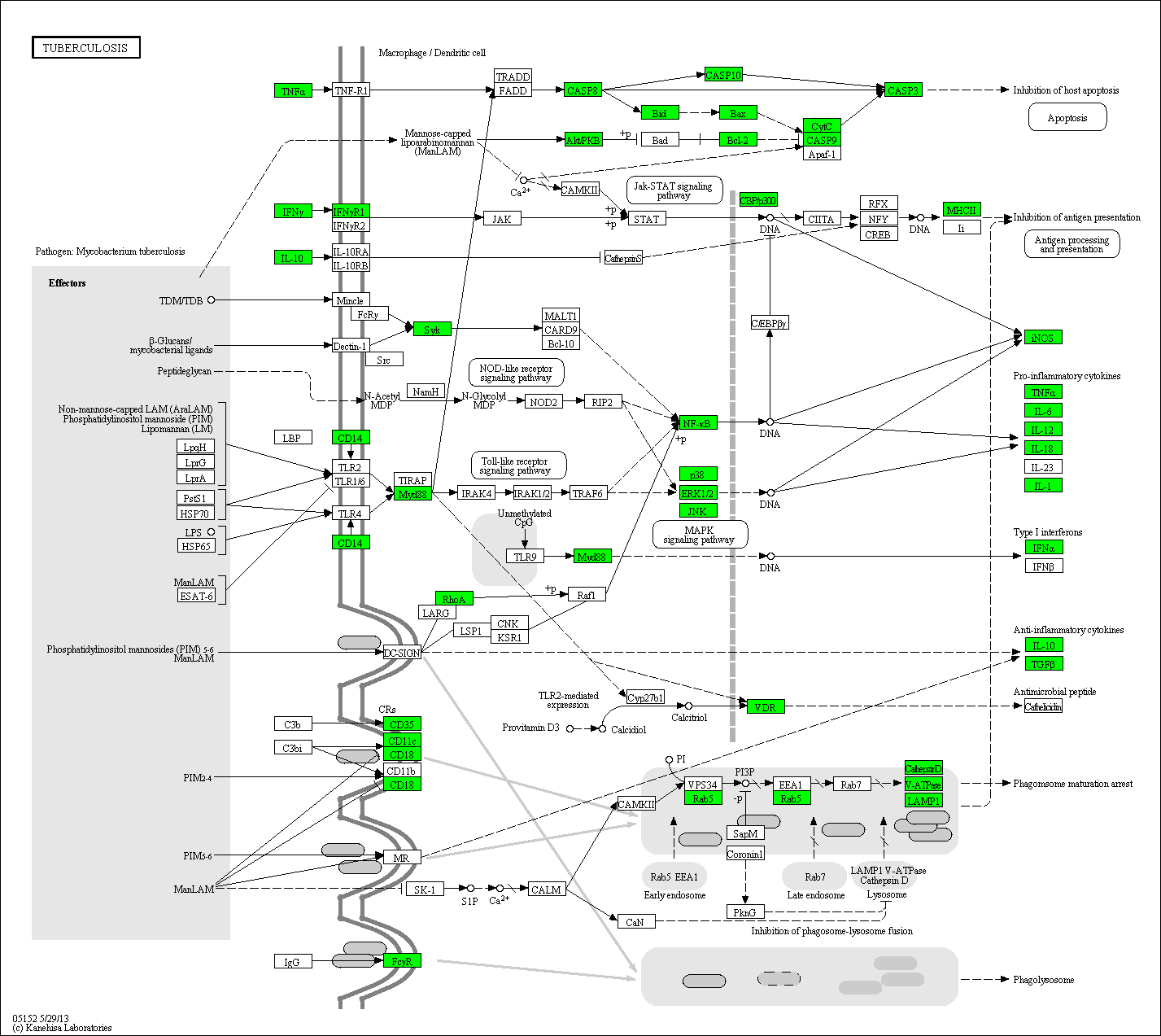
Proteins involved in Tuberculosis
Fig. 9: Pathways involved in Tuberculosis are shown. Proteins covered by the scioDiscover service are shown in green.